Quantifying the genetic differentiation among populations from molecular marker data play a pivotal role in quantitative genetics, conservation genetics and many areas of evolution and ecology.
The widely applied genetic differentiation statistics FST and GST have recently been criticised for underestimating differentiation when applied to highly polymorphic markers such as microsatellites.
Wang J. 2015. Does GST underestimate genetic differentiation from marker data? Molecular Ecology 24: 3546-3558.
My study shows that GST gives accurate estimates and underestimates of differentiation when demographic factors are more and less important than mutations, respectively. In the former case, all markers, regardless of diversity (HS), have the same GST value in expectation and thus give replicated estimates of differentiation.
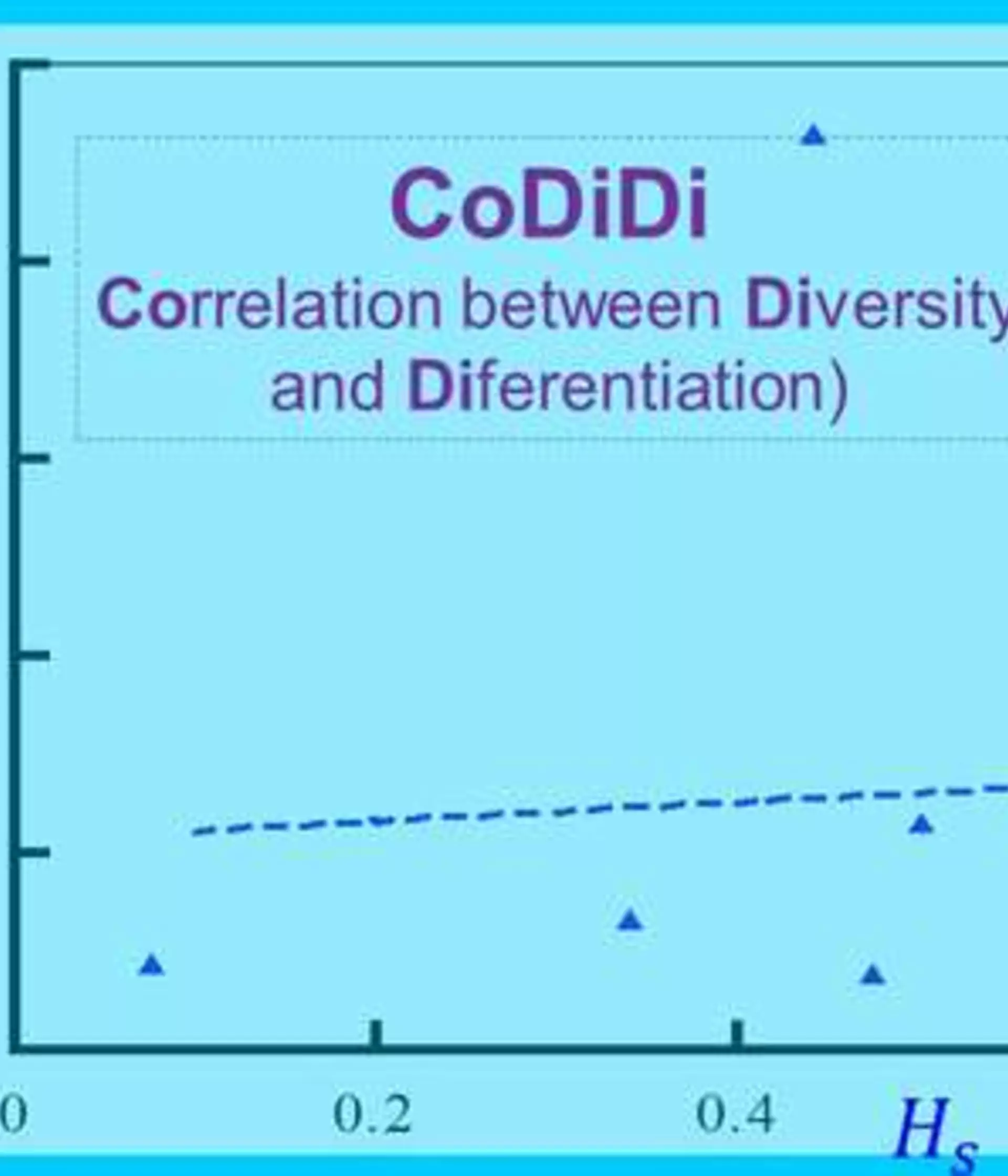
In the latter case, markers of higher HS have lower GST values, resulting in a negative, roughly linear correlation between GST and HS across loci. I propose that the correlation coefficient between GST and HS across loci, rGH, can be used to distinguish the two cases and to detect mutational effects on GST.
A highly negative and significant rGH, when coupled with highly variable GST values among loci, would reveal that marker GST values are affected substantially by mutations and marker diversity, underestimate population differentiation, and are not comparable among studies, species and markers.
The computer program, CoDiDi (Correlation between Diversity and Diferentiation), was created in year 2015 to calculate single locus GST and HS values, to test whether a single locus GST value is significantly different from 0 or not by permutations, and to calculate and test the significance of the correlation between GST and HS. The downloaded package includes user’s manual, example datasets, source code of the program in Fortran 90, and executables runnable on MS-DOS (Windows) and x-terminals (Max or linux).