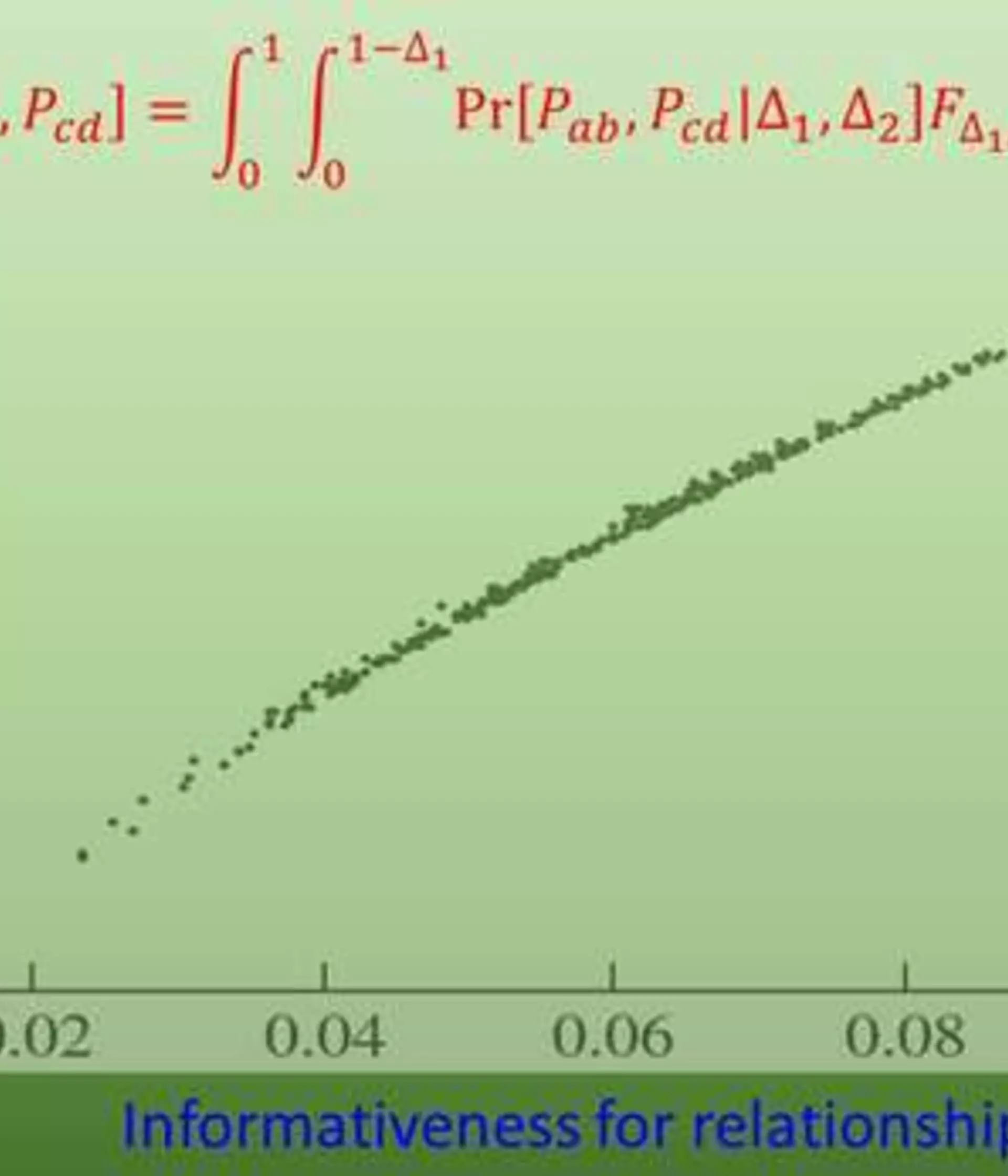
KinInfor (Kinship Informativeness) is a Fortran program that calculates the informativeness of markers in inferring pairwise relatedness or relationships.
Measuring the information content of markers in relationship/relatedness inferences is important in selecting highly informative markers to attain a given statistical power with the minimal genotyping effort.
This program calculates four informativeness measurements, IR, Ir, RMSD and PWR, for each single marker, and the last two measurements for multiple loci.
All four measurements incorporate genotyping errors and mutations, and are applicable to codominant and dominant genetic markers and to diploid and haploid individuals. Details of the four informativeness measurements are described in:
Wang, J (2006) Informativeness of genetic markers for pairwise relationship and relatedness inference. Theoretical Population Biology 70: 300-321.
The KinInfor program was created (version 1) in year 2006, and was updated (current version 2) in year 2016. The downloaded package includes source code in Fortran 90, an example dataset, the user’s manual, and the program executable runnable on MS-DOS in Windows 10. Mac or linux users need to generate the binary by compiling the source code and linking to NAG.